The Drosophila split-GAL4 system, first described in Luan et al., 2006, allows for restricted expression of regulatory targets only in cells where the two components of the split-GAL4 activator are co-expressed. As shown below, the GAL4 DNA-binding domain fused to the Zip- protein-pairing domain can be expressed in one pattern, a transcriptional activation domain fused to the Zip+ protein-pairing domain can be expressed in another pattern and the UAS reporter construct will be expressed in the intersection of the two patterns. We now have thousands of stocks that carry either 1) a construct that expresses a DNA-binding domain protein derived from either GAL4 or lexA or 2) a construct that expresses a transcriptional activation domain derived from either GAL4, VP16 or p65.
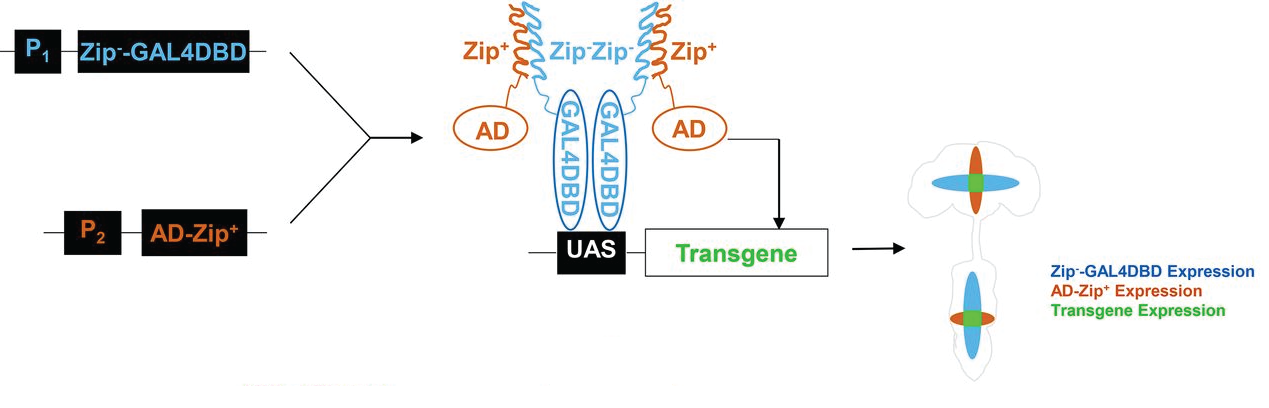
Figure adapted from Dolan et al., 2017
Split-GAL4 driver expression patterns: The majority of our split-GAL4 lines were generated at either the Janelia Research Campus (donated by Gerry Rubin) or at the Institute of Molecular Pathology in Vienna (donated by Barry Dickson). Information on the expression profiles of these lines can be found on the GAL4 DBD and p65 AD pages.
Hemidrivers
Stocks carrying split-GAL4 or split-lexA hemidrivers with zip adaptors:
- GAL4 DNA binding domain
- p65 activation domain
- GAL4 activation domain
- lexA DNA binding domain
- VP16 activation domain
Stocks carrying split-GAL4 intein hemidrivers
- Split-intein hemidrivers and combinations (note, these are incompatible with the other stocks listed on this page)
Tools
- Killer Zipper can be used to further restrict expression by including a construct expressing a protein that competes with the activation domain for binding.
Other
- Split-GAL4 or split-lexA combination stocks
- Transcription-based calcium sensors (split components bound to calmodulin or a calmodulin target)
- Shine-GAL4 (light-induced split-GAL4s)

