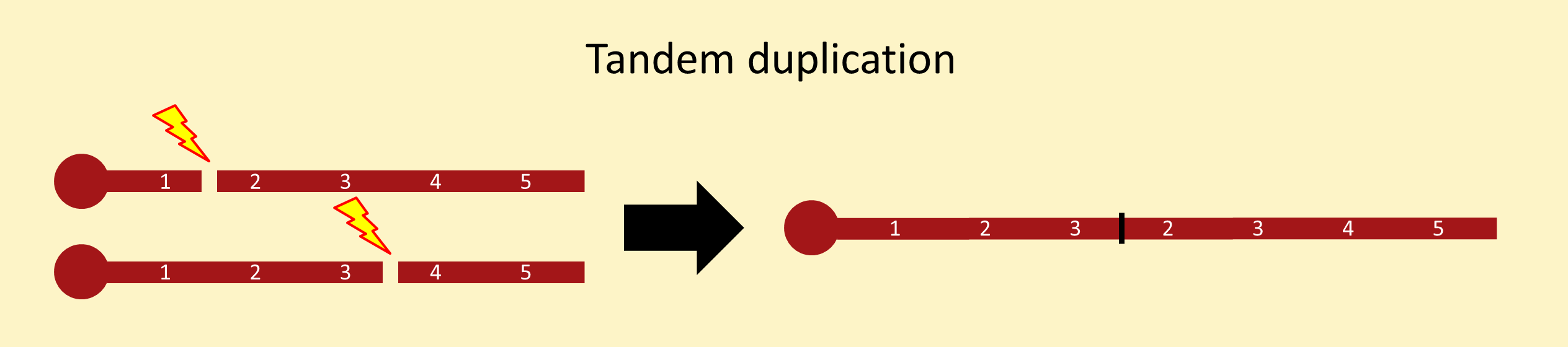
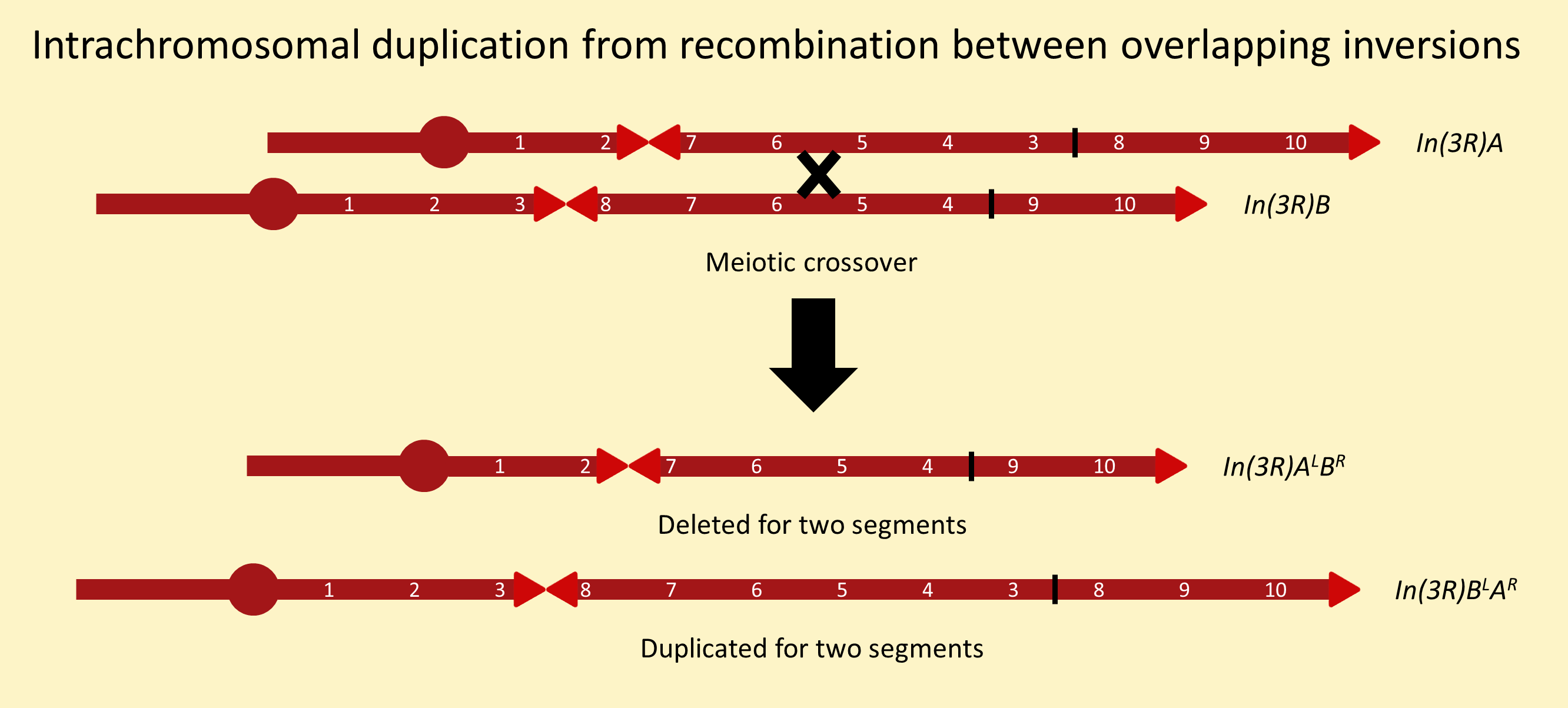
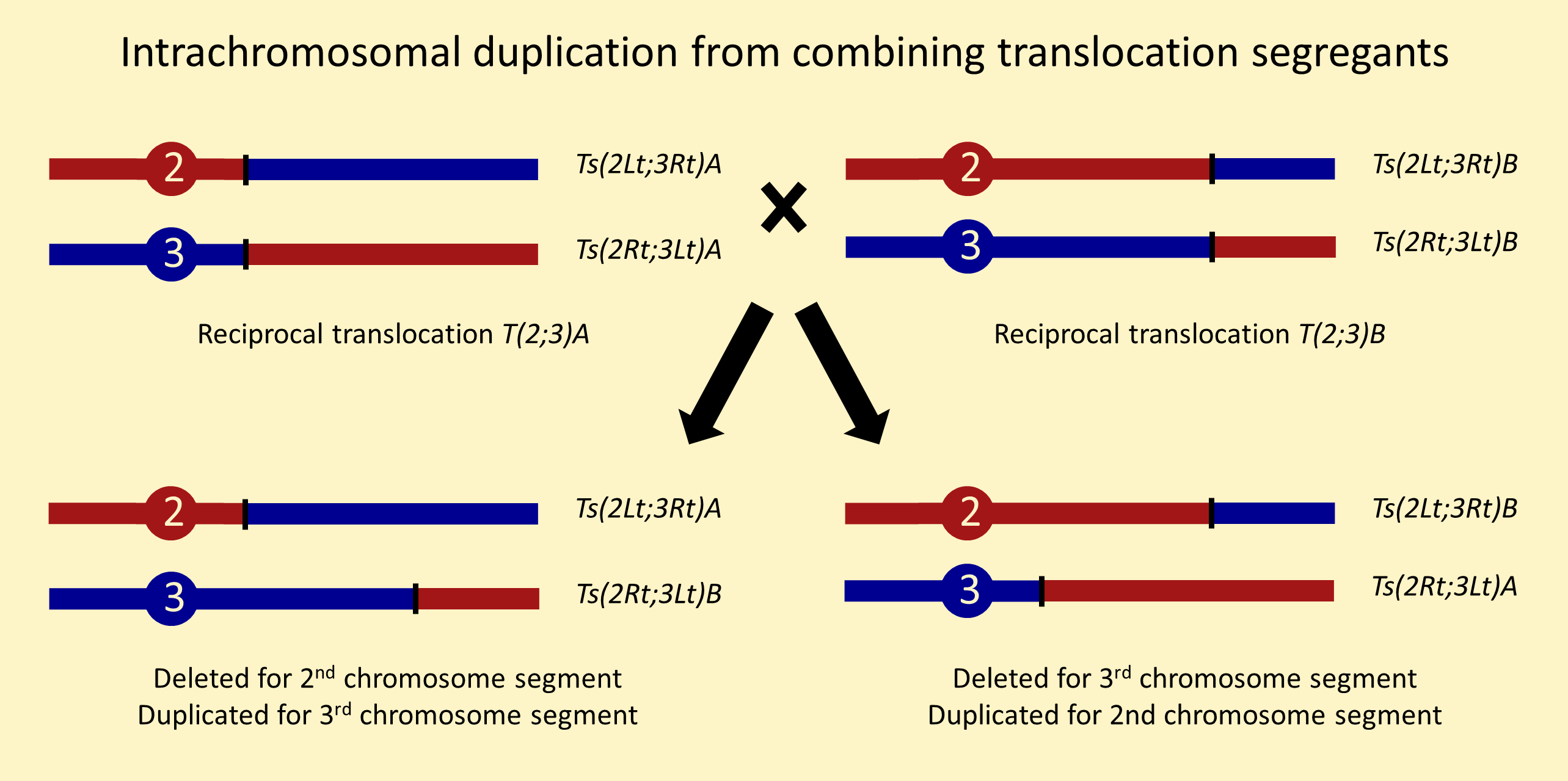
There are many ways to recover chromosomes carrying internally duplicated segments. The methods below are numbered to match the intrachromosomal duplications stock page. For a more complete review of duplications, see chapter 15 of Drosophila: a laboratory handbook.
1. Transposition
The most straightforward method to recover chromosomes carrying an extra copy of a segment is transposition, where three chromosome breakpoints allow the movement of a segment to a new insertion site as shown here.
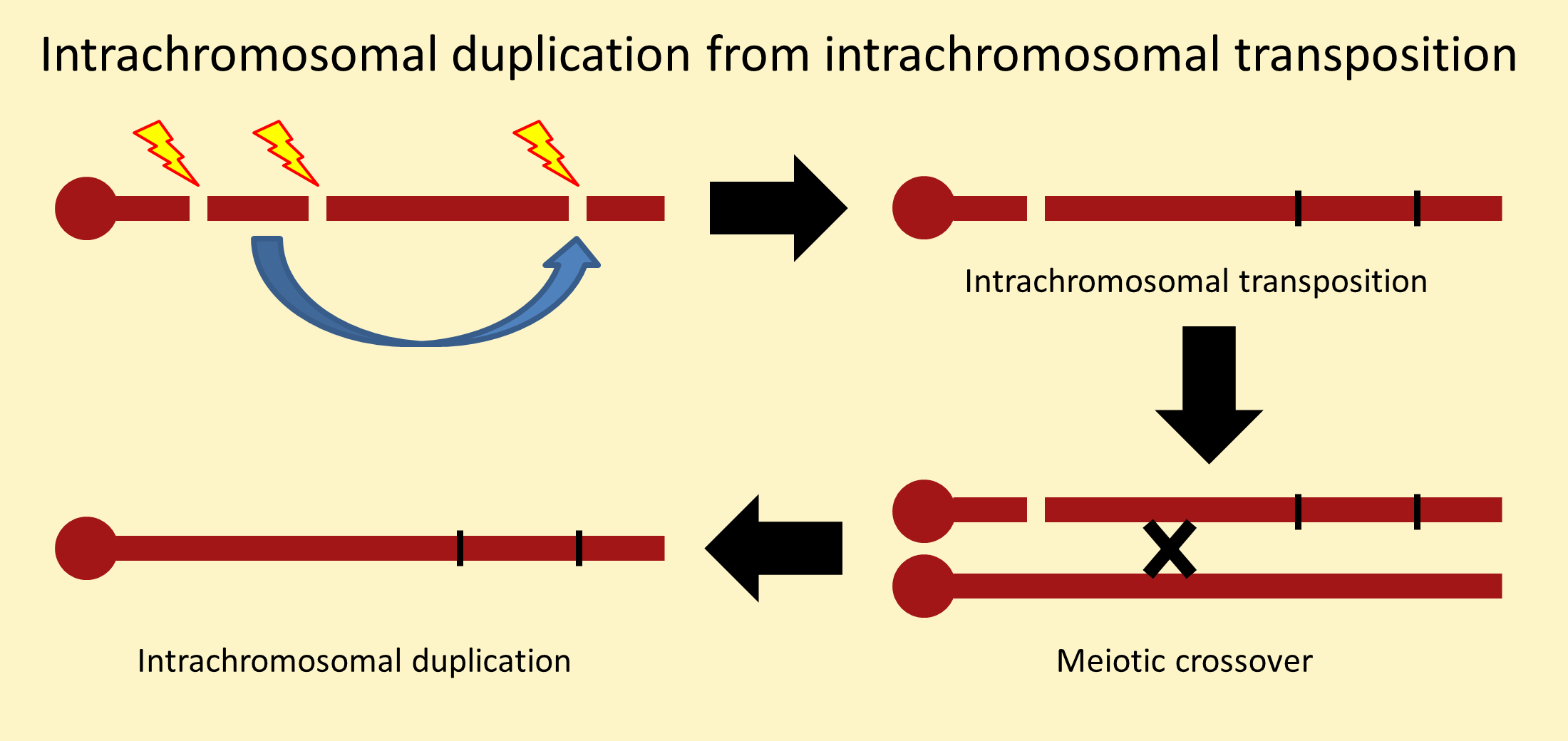
It is important to note the distinction between a transposition and a duplication. Though often used interchangeably, a transposition (Tp) consists of a deletion (Df) and a duplication (Dp). The duplication component of an intrachromosomal transposition can be separated from the deletion component by meiotic recombination. Intrachromosomal duplications are denoted Dp(1;1), Dp(2;2), etc.